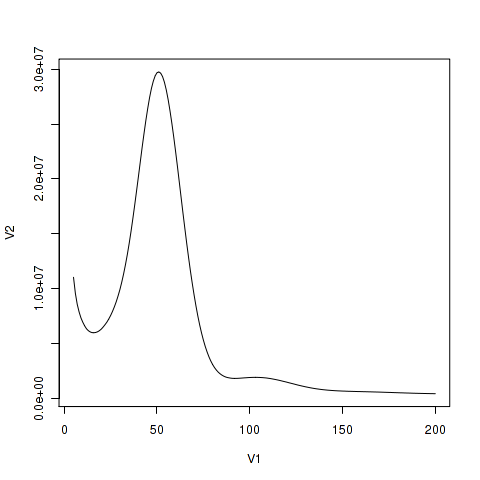
K-mer analysis and genome size estimate

Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies, Genome Biology

Allele-aware chromosome-level genome assembly of the autohexaploid Diospyros kaki Thunb

Plants, Free Full-Text

K-mer (k = 17) analysis for estimating the genome size of R.

19 K-mer analysis for estimating the genome size of Schizothorax
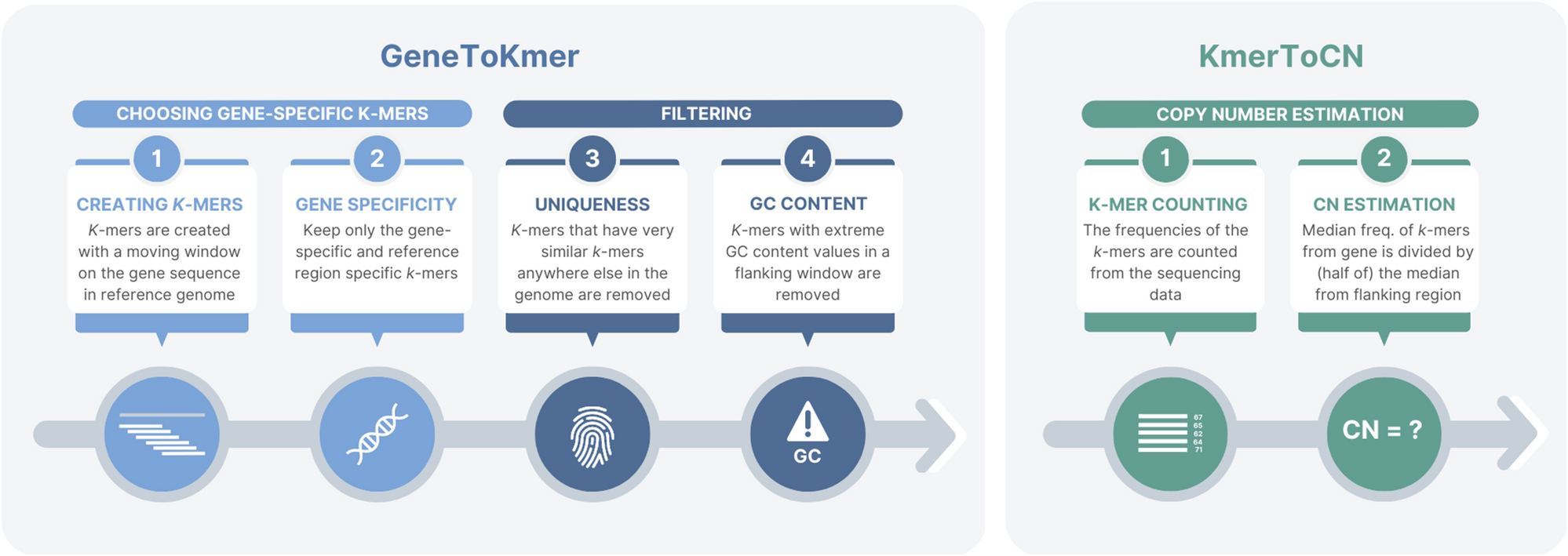
GeneToCN: an alignment-free method for gene copy number estimation directly from next-generation sequencing reads

A beginner's guide to assembling a draft genome and analyzing structural variants with long-read sequencing technologies - ScienceDirect

K-mer (k = 21) spectrum analysis to estimate genome size in the

Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies, Genome Biology
Large-scale k-mer-based analysis of the informational properties of genomes, comparative genomics and taxonomy

K-Mer-Based Genome Size Estimation in Theory and Practice

Measuring genome sizes using read-depth, k-mers, and flow cytometry: methodological comparisons in beetles (Coleoptera)
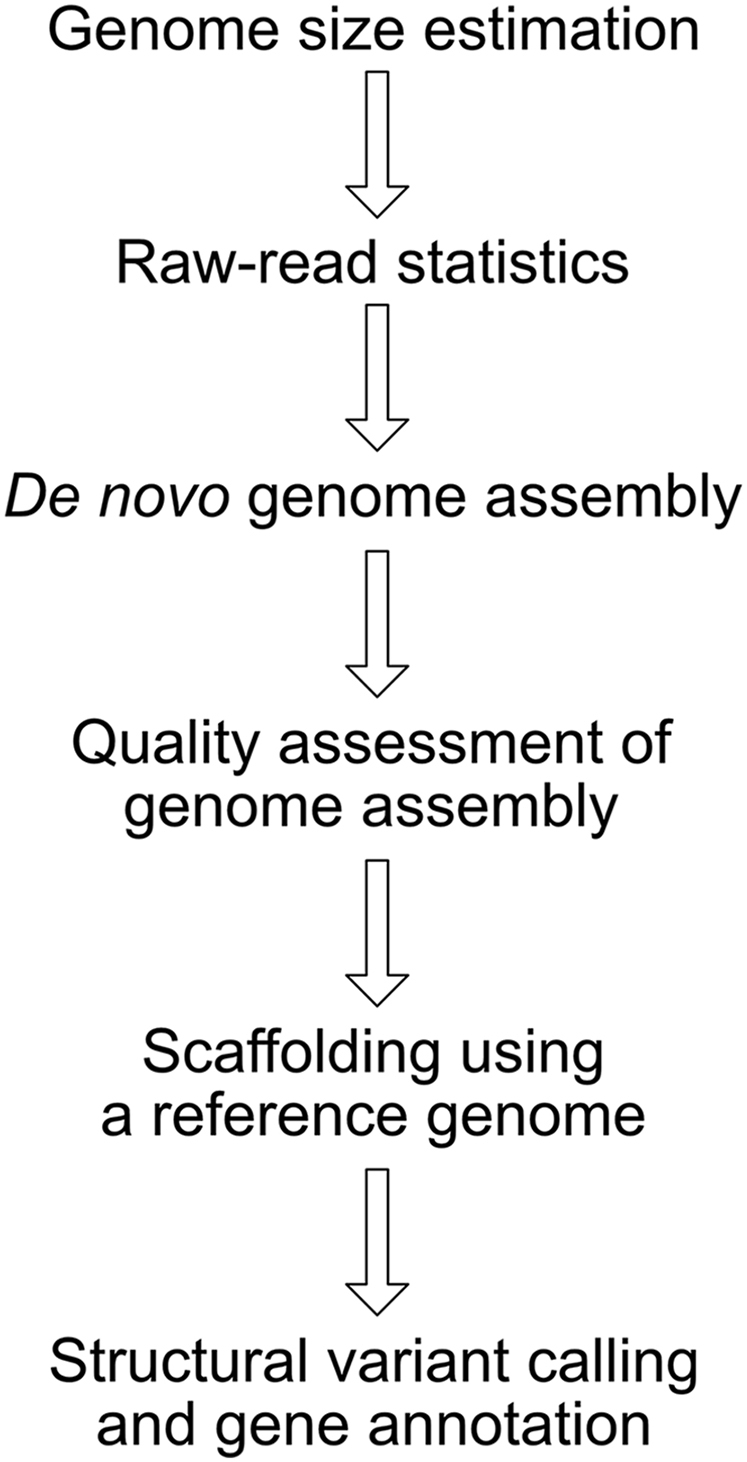
A beginner's guide to assembling a draft genome and analyzing structural variants with long-read sequencing technologies