PDF) SPANDx: A genomics pipeline for comparative analysis of large

Comparing different pipelines on the UHRR Iso-Seq Dataset. a

PDF) SPANDx: A genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets

CloVR-Comparative

PDF) Comparative genomic analysis identifies X-factor (haemin)-independent Haemophilus haemolyticus: a formal re-classification of 'Haemophilus intermedius

Can non-typeable Haemophilus influenzae carriage surveillance data

A technology-agnostic long-read analysis pipeline for
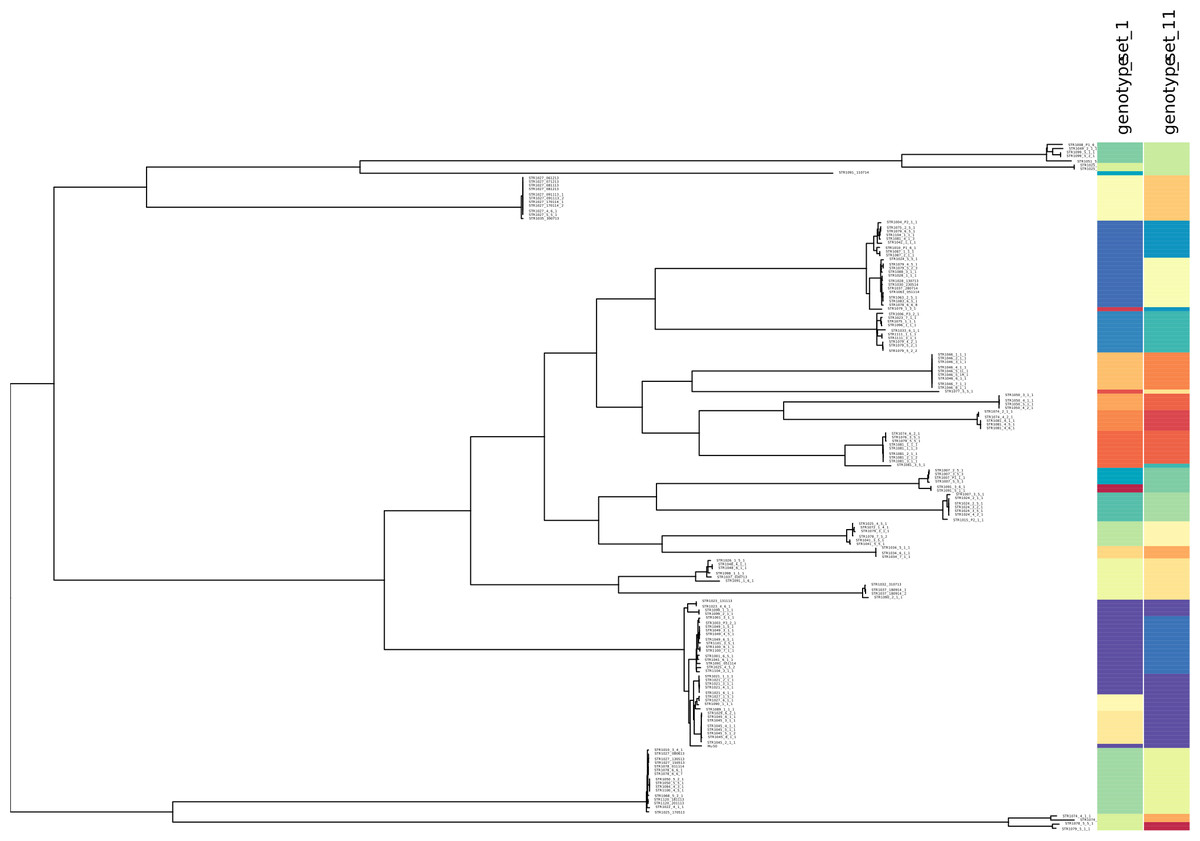
minSNPs: an R package for the derivation of resolution-optimised

Comparative analysis of whole-genome sequencing pipelines to

PDF) SPANDx: A genomics pipeline for comparative analysis of large haploid whole genome re-sequencing datasets

Comparative genomics of Nocardia seriolae reveals recent

Standardized phylogenetic and molecular evolutionary analysis

PDF] The Genome Analysis Toolkit: a MapReduce framework for

PDF) SPANDx: A genomics pipeline for comparative analysis of large
Longitudinal whole-genome based comparison of carriage and








